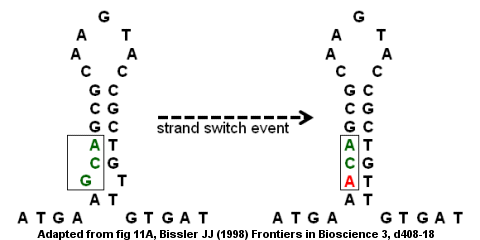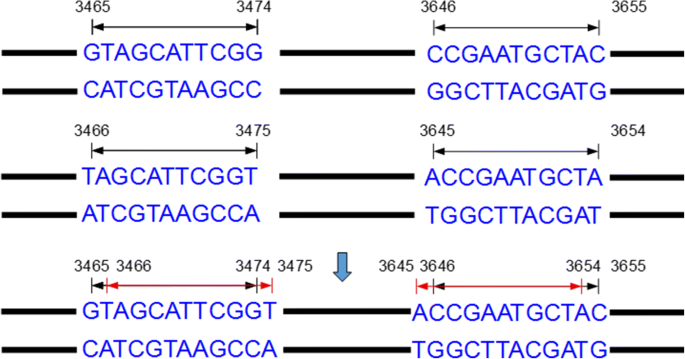
MiteFinderII: a novel tool to identify miniature inverted-repeat transposable elements hidden in eukaryotic genomes | BMC Medical Genomics | Full Text
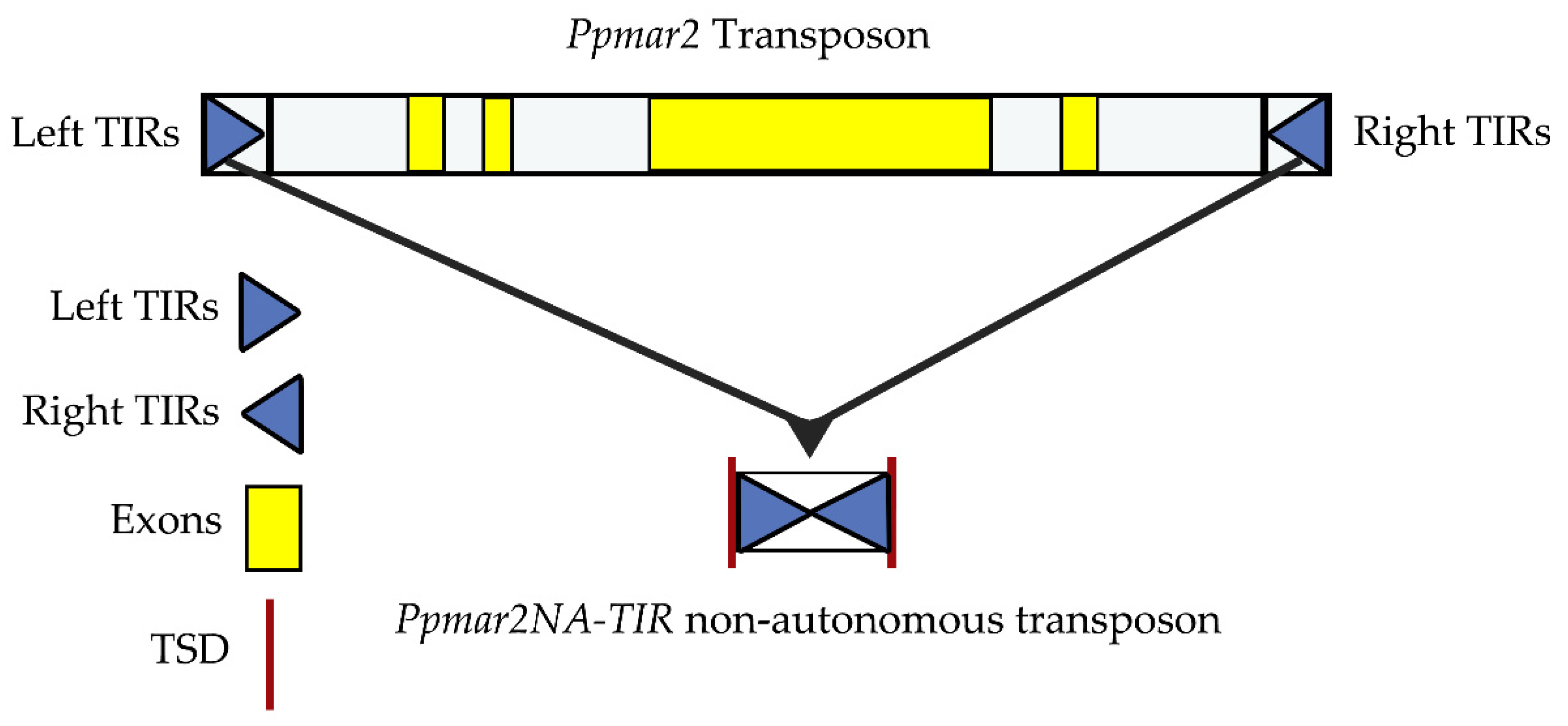
IJMS | Free Full-Text | Affinities of Terminal Inverted Repeats to DNA Binding Domain of Transposase Affect the Transposition Activity of Bamboo Ppmar2 Mariner-Like Element
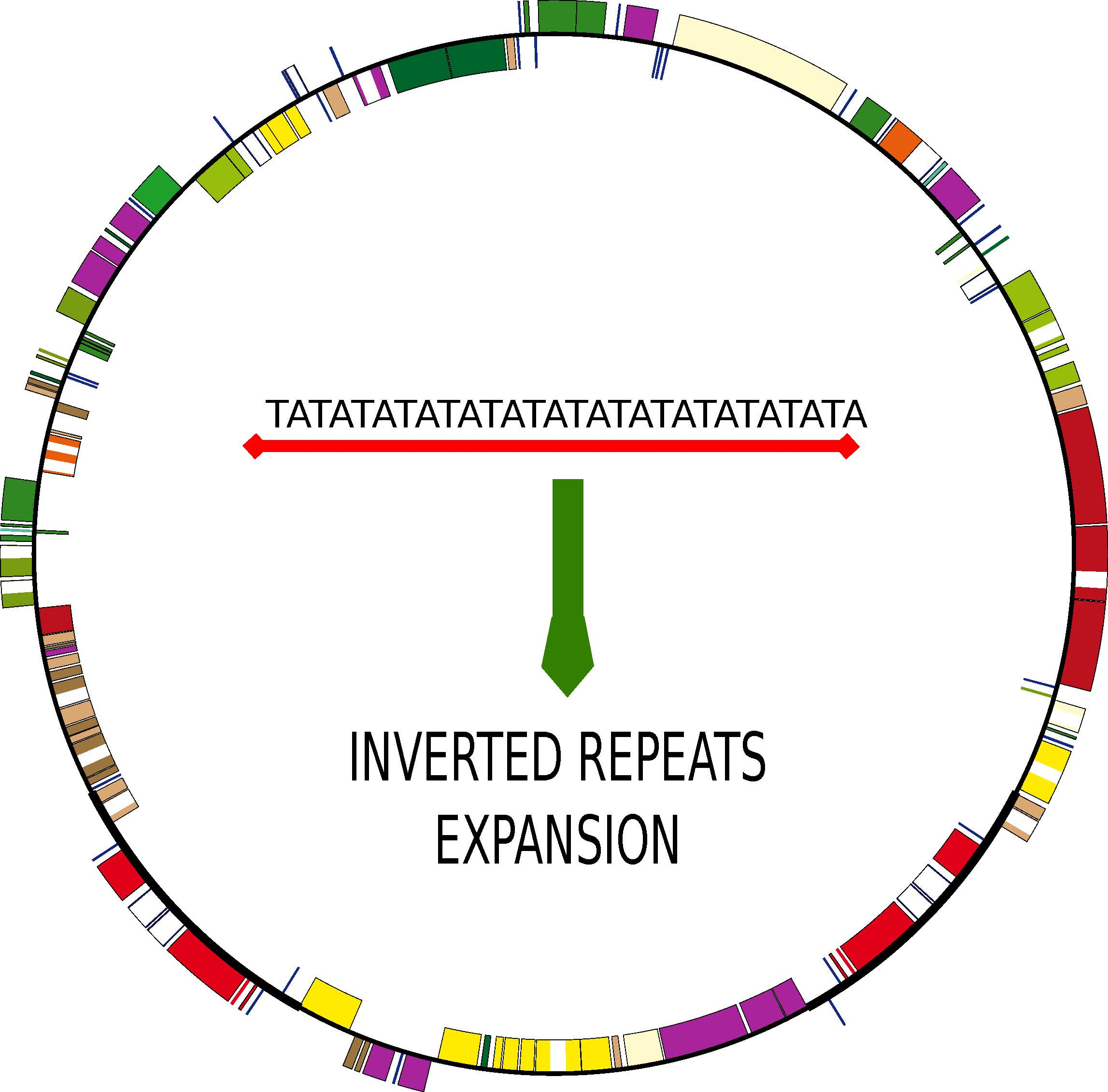
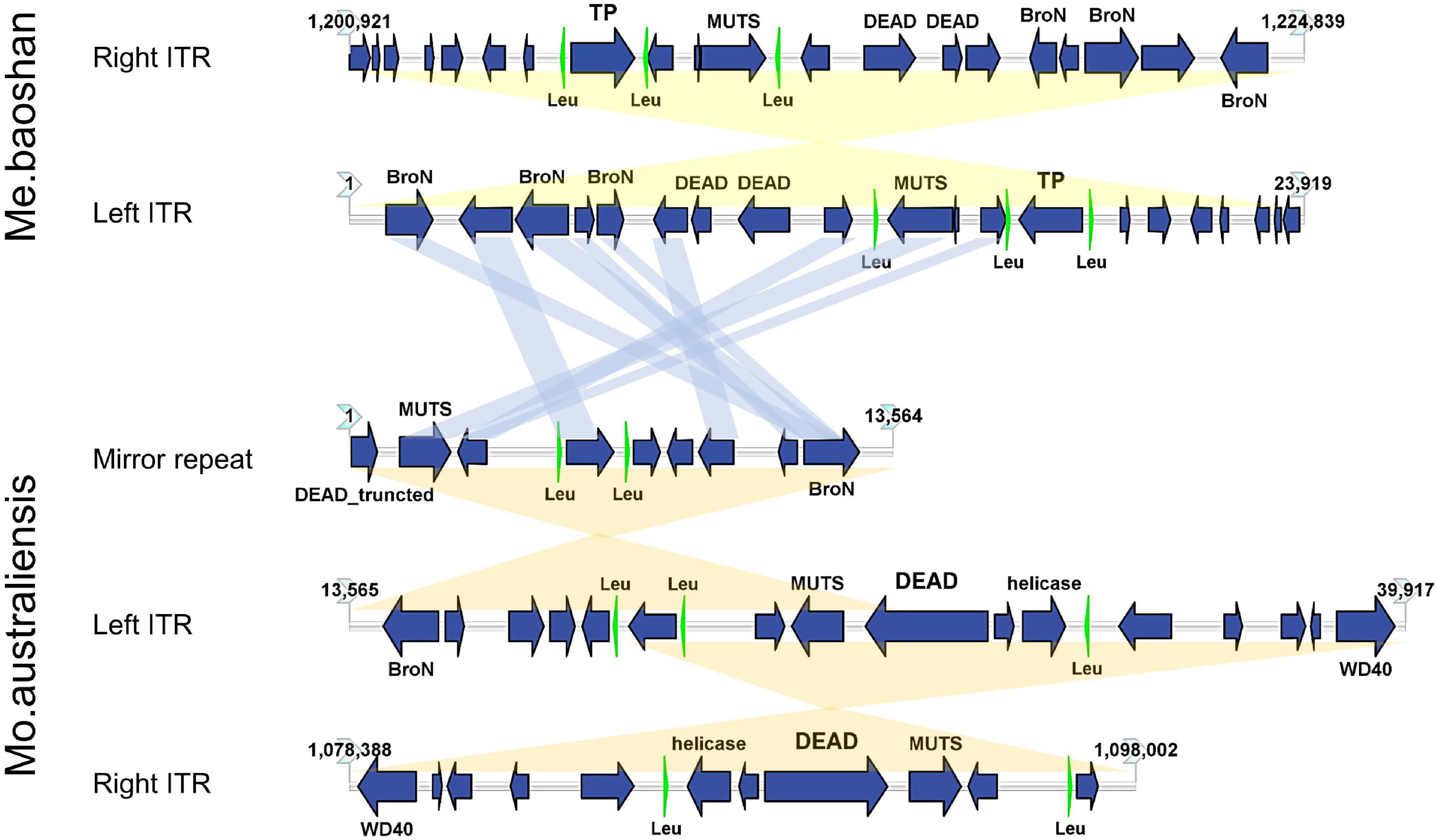
Genes | Free Full-Text | The Increase of Simple Sequence Repeats during Diversification of Marchantiidae, An Early Land Plant Lineage, Leads to the First Known Expansion of Inverted Repeats in the Evolutionarily-Stable
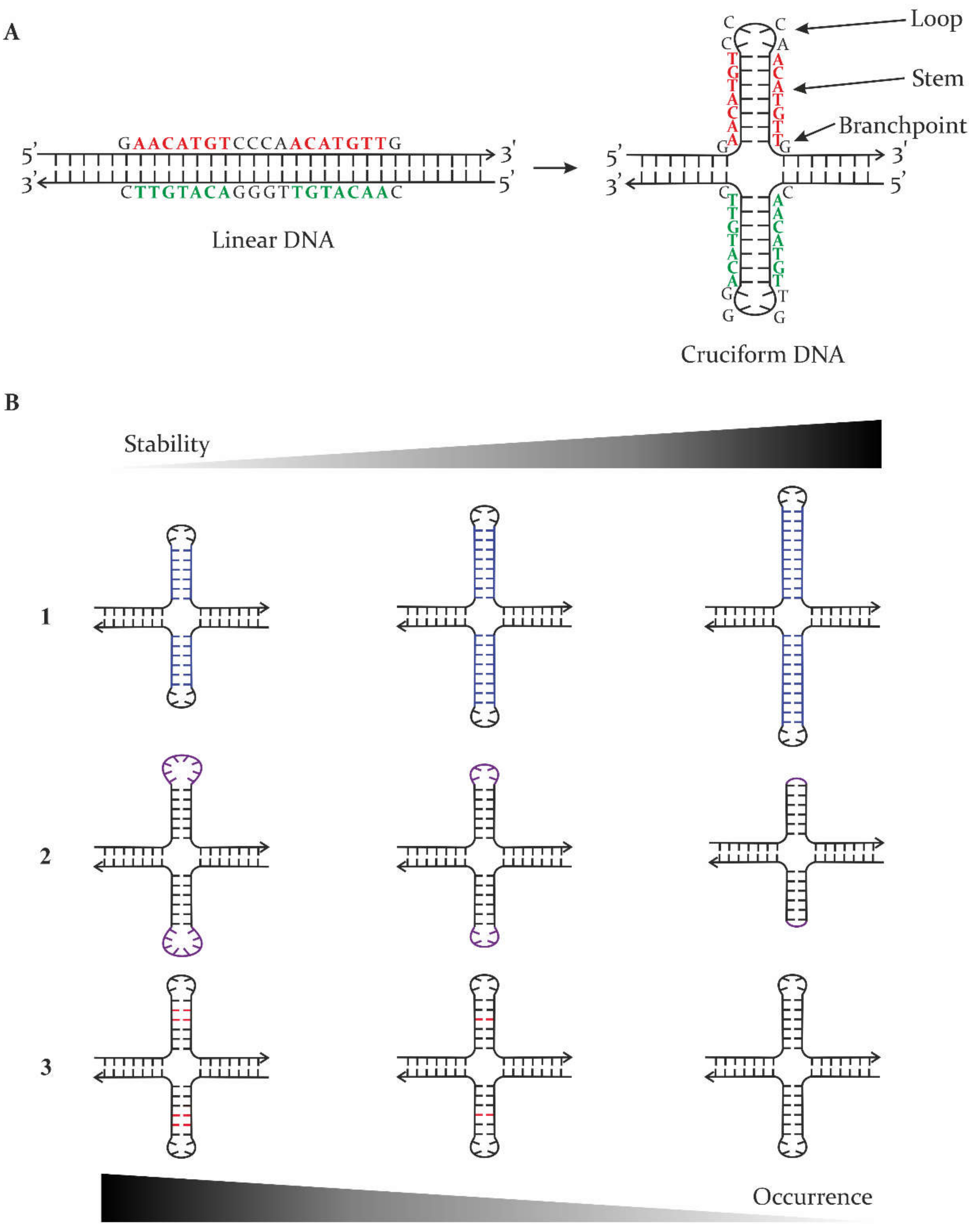
IJMS | Free Full-Text | Interaction of Proteins with Inverted Repeats and Cruciform Structures in Nucleic Acids
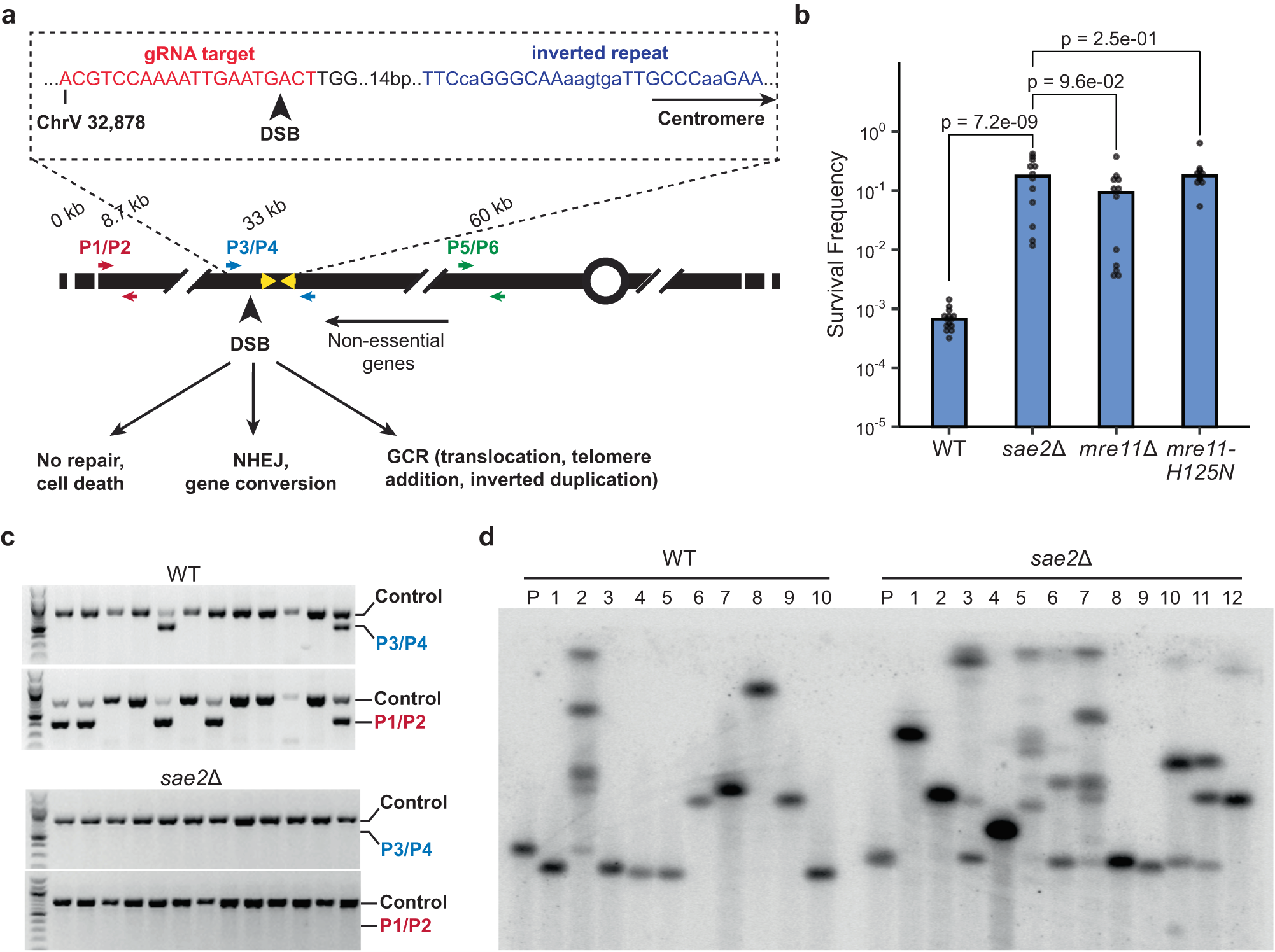
Double-strand breaks induce inverted duplication chromosome rearrangements by a DNA polymerase δ-dependent mechanism | Nature Communications
![PDF] Insertion Sequence Inversions Mediated by Ectopic Recombination between Terminal Inverted Repeats | Semantic Scholar PDF] Insertion Sequence Inversions Mediated by Ectopic Recombination between Terminal Inverted Repeats | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/f6d2ad1df500ba57e0b52b0ea614feb9f645c057/4-Figure4-1.png)
PDF] Insertion Sequence Inversions Mediated by Ectopic Recombination between Terminal Inverted Repeats | Semantic Scholar

Key steps in the origin-dependent inverted-repeat amplification model... | Download Scientific Diagram

MiteFinderII: a novel tool to identify miniature inverted-repeat transposable elements hidden in eukaryotic genomes | BMC Medical Genomics | Full Text

Replication stalling at unstable inverted repeats: Interplay between DNA hairpins and fork stabilizing proteins | PNAS

Global analysis of inverted repeat sequences in human gene promoters reveals their non-random distribution and association with specific biological pathways - ScienceDirect