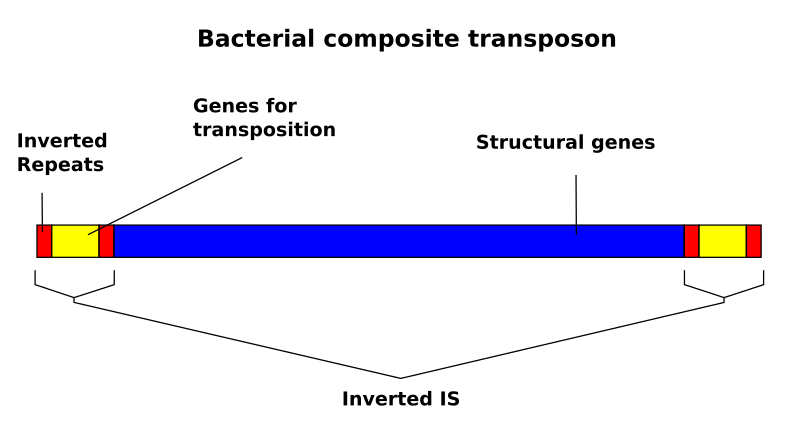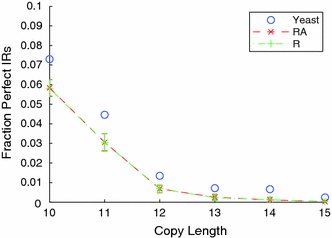
The distribution of inverted repeat sequences in the Saccharomyces cerevisiae genome | Current Genetics

MiteFinderII: a novel tool to identify miniature inverted-repeat transposable elements hidden in eukaryotic genomes | BMC Medical Genomics | Full Text
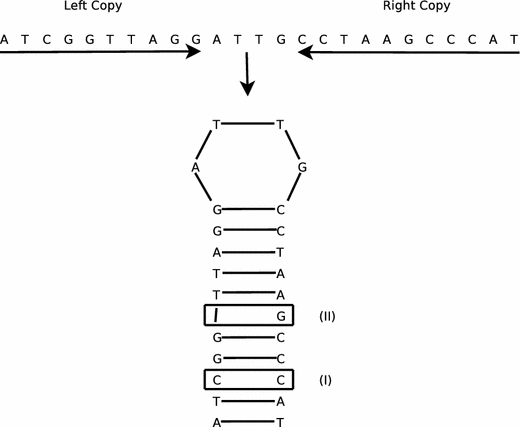
The distribution of inverted repeat sequences in the Saccharomyces cerevisiae genome | Current Genetics
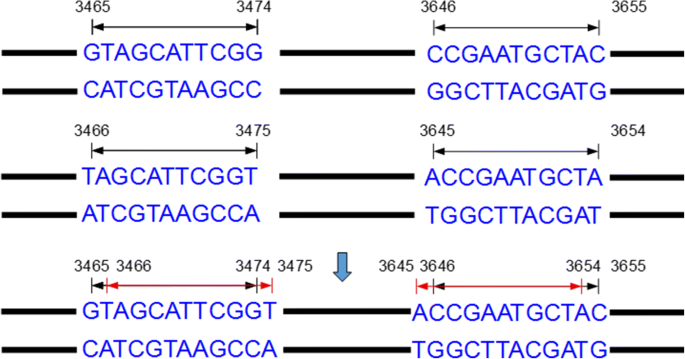
MiteFinderII: a novel tool to identify miniature inverted-repeat transposable elements hidden in eukaryotic genomes | BMC Medical Genomics | Full Text
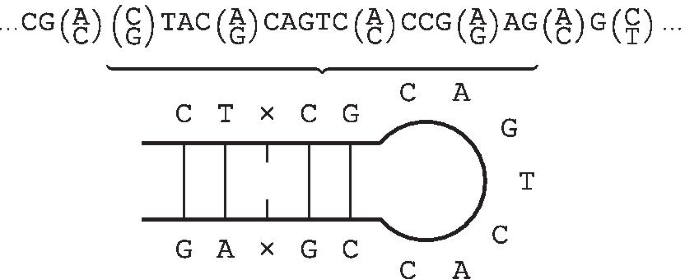
IUPACpal: efficient identification of inverted repeats in IUPAC-encoded DNA sequences | BMC Bioinformatics | Full Text

Expanded inverted repeat region with large scale inversion in the first complete plastid genome sequence of Plantago ovata | Scientific Reports

MUST: A system for identification of miniature inverted-repeat transposable elements and applications to Anabaena variabilis and Haloquadratum walsbyi - ScienceDirect

npInv: accurate detection and genotyping of inversions using long read sub-alignment | BMC Bioinformatics | Full Text

His 6 -BrxR binds DNA inverted repeats in vitro. (A) Sequence of pEFER... | Download Scientific Diagram

The distribution of inverted repeat sequences in the Saccharomyces cerevisiae genome | Current Genetics